Introduction
Materials and Methods
Seed material, test conditions and sampling
RNA isolation and cDNA synthesis
Primer designing
RT-PCR analysis
In silico analysis of miRNA and cis-acting elements
Results
Shortlisted genes for expression studies
Genes induced in SI and GN1 genotypes
Genes induced in GN1 and GN3 genotypes
Genes induced in all three genotypes
Genes induced only in a specific genotype
Identification of miRNAs
Prediction of cis-acting elements and gene structure
Discussion
Introduction
Germination as a process represents the initial developmental stage in the life cycle of a plant, marking the onset of a new generation, and its successful execution is a prerequisite for initiating this cycle (Bewley, 1997). Germination encompasses the sequence of occurrences that begin with the absorption of water by the dormant, dehydrated seed and culminate in the elongation of the embryonic axis. The process of seed germination presents a formidable challenge in reviving dormant seeds and transitioning them into active seedlings (Baskin et al., 2019). Several environmental factors, including temperature, soil moisture, light, and nutrient availability, exert varying influences on this process, which differ across genotypes. The delicate balance between the two phytohormones, gibberellins (GAs) and abscisic acid (ABA), assumes a significant role in the regulation of seed germination. ABA induces dormancy, while GA promotes germination, and mutants affecting the levels or signaling pathways of these hormones frequently manifest as seed dormancy phenotypes (Holdsworth et al., 2008; Ji et al., 2022; Kim et al., 2023a, 2023b; Kim et al., 2024). Dormancy is favored when the ABA/GA ratio is high, whereas germination is promoted when this ratio is low (Holdsworth et al., 2008). Additionally, other phytohormones, such as ethylene, interact with ABA and influence dormancy and germination processes by triggering endosperm rupture (Linkies et al., 2009). Notably, genes associated with GA and ABA biosynthesis, perception, and signaling were among the first loci identified in Arabidopsis mutants that are associated with dormancy and germination (Holdsworth et al., 2008).
Seed dormancy, which refers to the temporary inability of seeds to germinate even when all the necessary conditions for germination are met, is a common trait in wild plants that serves as a survival mechanism under unfavorable environmental conditions (Bewley, 1997). While the incorporation of native plants into agriculture has often been accompanied by selection against seed dormancy, many modern crops still carry genes associated with this trait. In most cases, seed dormancy is not absolute, which means that not all seeds in a population are dormant or non-dormant, and it is often expressed quantitatively. For instance, in cucurbit crops, it is common to encounter seed lots with delayed germination, partial dormancy (Nerson, 2002), or short-term dormancy (Odland, 1937), where seed germination improves with increased storage compared to germination at harvest. Partial and short-term dormancy are more likely to occur in immature seeds compared to fully mature seeds. Genetic regulation of seed dormancy in cucumber was studied using different cultivars, including the dormant cultivar 'Boroda' and the non-dormant cultivar 'Marketer', as described in a study conducted by Ali et al. (1991). Another study by Amritphale et al. (1993) reported the induction of secondary dormancy in cucumber through exposure to two short cycles of red-far red light, lasting 15-60 minutes, during seed incubation.
Seed dormancy can be overcome by imbibition of seeds at low temperatures, also known as stratification, or by dry storage for a certain period, known as after-ripening. Germination, which involves the radicle protruding through the surrounding structures such as the endosperm and testa in Arabidopsis, can take place when non-dormant seeds are exposed to favorable environmental conditions in terms of humidity, light, and temperature (Finch-Savage et al., 2006).
Watermelon (Citrullus lanatus) is a valuable and economically significant fruit crop, extensively grown in China, Turkey, India, Brazil, and Algeria, with a staggering annual production of more than 132 million tons in 2021 (FAOSTAT, 2021). Numerous studies have focused on improving the nutritional quality, sweetness, graft survival rate, and tolerance to biotic and abiotic stresses in watermelon (Dabirian and Miles, 2017; Li et al., 2017; Zhang et al., 2020). However, there has been limited research on watermelon seed germination. Dormant watermelon seeds can be germinated by clipping the seed apex. Triploid watermelon seeds are smaller in size compared to diploid varieties, resulting in limited reserves for germination and seedling growth (Wu et al., 2022). Efficient and consistent seed germination is vital in commercial agriculture to optimize the utilization of production resources. Hence, enhancing the rate and quality of watermelon seed germination is crucial for promoting healthy growth of seedlings and early availability in the market. Therefore, in this study, we aimed to characterize the genes involved in germination and dormancy in watermelon.
In this study, we characterized 25 genes related to germination and dormancy in watermelon. We identified four genes which could be involve in the germination and two genes that are involved in dormancy. We also identified some miRNA and cis-acting elements that may be regulating gene expression related to germination or dormancy in watermelon.
Materials and Methods
Seed material, test conditions and sampling
Three cultivars were selected for this study based on their germination percentage: the SI cultivar with good germination of 96%, the GN1 cultivar with moderate germination of 52% and GN3, with low germination of 4% under optimal germination conditions (Fig. 1). Around 50 seeds of each cultivar were imbibed for 24 hours in a muslin cloth. After imbibition, the seeds were taken and excess water was removed using autoclaved tissues. These seeds were then kept in a sterile petri dish containing sterile filter paper moistened with sterile water. The moisture was maintained throughout the experiment using sterile water. These seeds were maintained in an incubator at 25°C. Five seeds at each time point were sampled at 0h, 6h, 12h, 24h and 36h. Samples were stored in -80°C until further use.
RNA isolation and cDNA synthesis
The seed samples were ground to powder with liquid nitrogen using a mortar and pestle and total RNA was extracted using TaKaRa MiniBEST Plant RNA Extraction Kit (TaKaRa, Japan) following the manufacturer’s guidelines. Three micrograms of RNA were used for first-strand cDNA synthesis using PrimeScript 1st strand cDNA Synthesis Kit (TaKaRa, Japan) following the manufacturer’s guidelines.
Primer designing
Twenty-five genes were selected based on previous reports. Gene sequences were downloaded from the Arabidopsis database (https: //www.Arabidopsis.org). Later, we performed a Blast search with default parameters of the watermelon genomics database (http://cucurbitgenomics.org/blast). Primers were designed in the CDS region using Primer3 tool software (https://primer3.ut.ee/). After designing, these primers were checked for specificity using the in silico PCR tool (http:// insilico.ehu.es/PCR/).
RT-PCR analysis
Real time PCR was performed in a LightCycler® instrument (Roche, Mannheim, Germany) using SYBR green chemistry. The watermelon ACTIN gene was used as the endogenous control. PCR reactions were carried out in 10 µL reaction containing 1 µL of first strand DNA, 5 pmol primers and 5 µL SYBR green mix (TaKaRa, Japan). The thermal profile used was: Pre-incubation at 95°C for 10 min followed by 45 cycles of 95°C for 20 s, annealing at 60°C for 15 s and final extension of 72°C for 30 s. Melt curve analysis was also performed to determine the specificity of the reaction. To calculate mean relative expression levels, cDNAs from three biological samples in two technical replications each were used. Relative expression was calculated using 2-∆∆Ct value (Livak and Schmittgen, 2001). The data were analyzed using the Roche inbuilt Software (Roche, Germany) with default baseline and threshold. Relative transcription levels are presented graphically. Results are presented as mean ± SE of relative expression in comparison with corresponding uninfested control sample (0 hrs). Mean values higher than 2-folds were compared through paired t test at P &5 level using Microsoft Excel program.
In silico analysis of miRNA and cis-acting elements
Shortlisted 25 genes were used to predict the gene structure using online tool GSDS2.0 (Gene Structure Display Server 2.0 (gao-lab.org) between Arabidopsis and watermelon. Similarly, to predict the putative miRNA in the coding region of 25 genes using the psRNATarget online tool (http://plant grn.noble.org/psRNATarget/analysis) was used. The putative cis-regulatory elements located within 1500 bp upstream from the transcription start point of 25 genes were analyzed using the Plant-Care database (http://bioinformatics.psb.ugent. be/webtools/plantcare/html/) (Lescot et al., 2002).
Results
Shortlisted genes for expression studies
Based on earlier reports on seed germination and dormancy in Arabidopsis, 25 genes were shortlisted for gene expression studies (Table 1). Out of the twenty-five genes, 11 genes, related genes or transcriptional factor that regulate the ABA signaling pathway, viz. ABA1, ABA2, ABA3, NCED6, ABI3, ABI4, ABI5, MDN1, CRU3, CyP707A1 and CYP707A2. Similarly, five genes from GA signaling pathways namely GA2ox2, GIDIB, SCL3, PKL and HD2B. Another two genes from ethylene pathway i.e, ACO1 and ACO4 were also selected. MFT gene may involve in the JA signaling, and another five genes were shortlisted.
Table 1.
List of genes and with designed primer pairs for RT-PCR analysis.
Genes induced in SI and GN1 genotypes
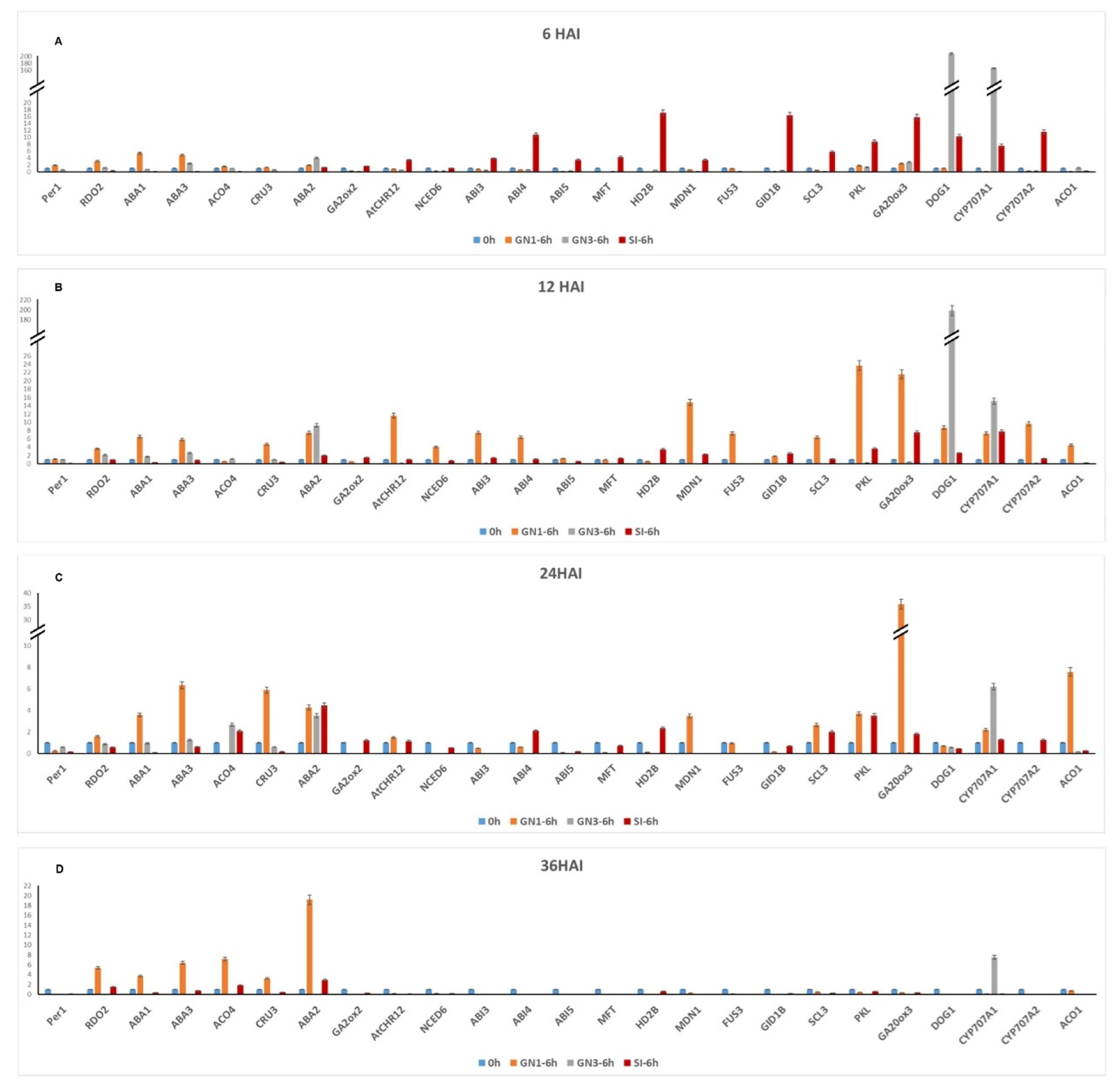
Out of the 25 genes tested, two genes ABI3 (Abscisic acid insensitive 3) and ABI4 (Abscisic acid insensitive 4) were up-regulated in GN1 at 12hai and SI at 6 and 12hai when compared to the control (0 hai). In addition, Midasin homolog 1 domain containing protein (MDN1) and Scarecrow like (SCL3) genes were up-regulated (≥ 2 fold) at both 12hai and 24hai in GN1; at 6hai and 12hai in SI genotype. Another two genes Chromatin‐remodeling (CHR12) and Cytochrome P450 (CYP707A2) were upregulated in GN1 genotype at 12hai and SI genotype at 6hai. However, Pickle (PKL) gene significantly upregulated in SI genotype at three time points (6, 12, 24hai) and in GN1 at 12 and 24hai (Fig. 2).
Genes induced in GN1 and GN3 genotypes
Transcription elongation factor tfiis/reduced dormancy 2 (RDO2) gene was up-regulated in GN1 genotype at 6hai and 12hai; GN3 genotype at 12hai and 36hai. Another gene, Abscisic acid deficient 3 (ABA3) was significantly up-regulated in all four time points in GN1 and only at 6hai and 12hai in GN3 genotype (Fig. 2).
Genes induced in all three genotypes
A total of five genes were expressed in all the three genotypes at different time points compared to their respective controls. Abscisic acid deficient 2 (ABA2) gene was up-regulated in GN1 at 12hai, 24hai, 36hai; in GN3 at 6hai, 12hai and 24hai; in SI at 12hai, 24hai and 36hai. In addition, GA 2-Oxidases (GA2ox3) gene was upregulated in GN1 at 6hai, 12hai and 24hai; GN3 at 6hai; SI at 6hai and 12hai. Another gene encoded 1-AMINOCYCLOPROPANE-1- CARBOXYLIC ACID (ACC) OXIDASE (ACO4) was induced at 36hai in GN1; 24hai in GN3 and at 24hai in SI genotype. The remaining two genes Delay of germination 1 (DOG1) gene upregulated in GN1 at 12hai; GN3 and SI at 6hai and 12hai. Cytochrome P450 (CYP707A1) gene was upregulated in GN1 at 12hai and 24hai; GN3 at 6hai, 12hai, 24hai and 36hai; SI at 6hai and 12hai respectively. However, the two genes DOG1 and CYP707A1, showed high expression of 150-200 fold change only in GN3 genotype (Fig. 2).
Genes induced only in a specific genotype
Out of the 25 genes, four genes were found to be up-regulated only in SI and six genes only in GN1 at different time points. Abscisic acid insensitive 5 (ABI5) and MOTHER-OF-FT (MFT) genes were upregulated at 6hai compared with their respective controls. However, HISTONE DEACETYLASE2B (HD2B) gene was induced at 6hai, 12hai and 24hai in SI; GIBBERELLIN INSENSITIVE DWARF 1 (G1D1B) was induced in SI at 6hai and 12hai. Further three genes namely 1-CYSTEINE PEROXIREDOXIN 1 (PER1) upregulated at 6hai, 9-CIS-EPOXYCAROTENOID DIOXYGENASES (NCED6) and FUSCA3 (FUS3) were upregulated at 12hai in GN1 genotype. Another gene 1-AMINOCYCLOPROPANE-1- CAR BOXYLIC ACID (ACC) OXIDASE (ACO1) was upregulated at 12 and 24hai; CRUCIFERIN3 (CRU3) gene was up-regulated at 12, 24 and 36hai respectively. However, ABSCISIC ACID DEFICIENT 1 (ABA1) gene was upregulated at all- time points in GN1 genotype (Fig. 2).
Identification of miRNAs
To identify miRNA present in the target genes all the transcript sequences were analyzed using the psRNA Target online tool. We have identified many miRNA target sites related to seed germination or seed dormancy in melon species in Per1, RDO2, CHR12, ABI3, ABI4, ABI5, MFT, MDN1, GIDIB, SCL3, PKL, GA20x3, CYP707A1 and CyP 707A2 genes. Remaining ten genes were not predicted with any miRNA target sites (Supplementary Table S1). Some miRNAs, such as miR-156, miR-396, miR-171, and miR- 390, were predicted in the multiple genes. Some were specific to only one gene, like miR-395 in PKL gene, miR-167 in GA20x3, and miR-166 in ABI5.
Prediction of cis-acting elements and gene structure
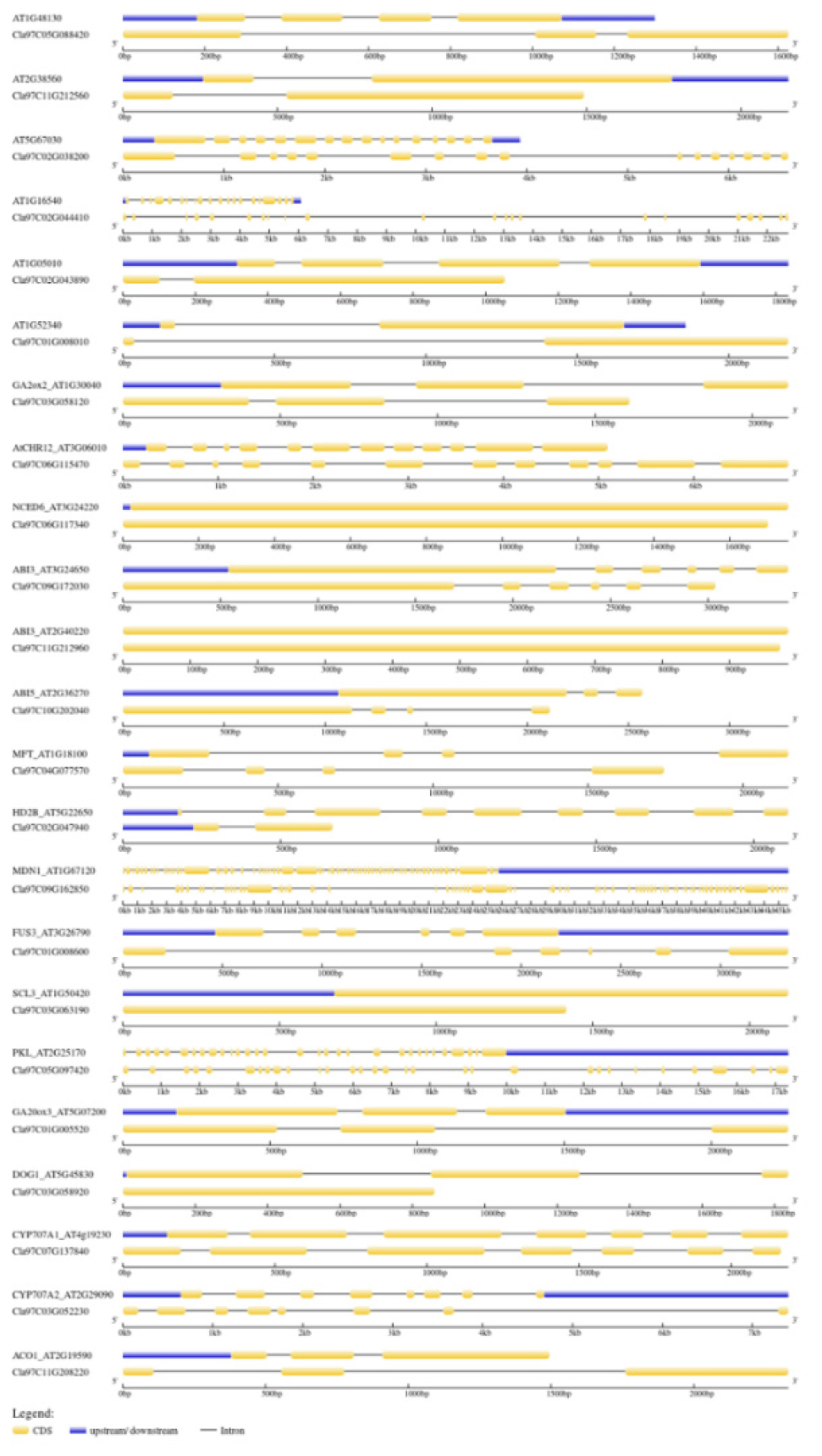
We have identified many cis‐regulatory elements related to phytohormones and seed germination/dormancy responses in the promoter regions of 25 target genes. Some of the cis-elements were commonly present in the most of the genes. These included ABA‐responsive elements; TGACG and CGTCA motifs; TAAT-box, G-box, CAAT-box, ARE, associated with the gibberellic acid response. Some of the cis-acting elements were specific to genes viz., AACA-motif and GCN4-motif detected in the ABI3 and PKL genes reported to be involved in the endosperm expression. Another cis-acting elements namely RY-element, involved in the regulation in the seeds present in the CRU3 gene. Analysis of the exon–intron organization revealed structural divergence between the watermelon and Arabidopsis were almost similar (Supplementary Table S2). The highest number of exons have been detected in MDN gene with 77 exons in watermelon and 74 exons in Arabidopsis followed by ABI3 with 22 and 20 in watermelon and Arabidopsis. The lowest exons were observed in the genes namely NCED6, ABI4 and SCL3 with only one exon in both watermelon and Arabidopsis (Fig. 3, Table 2).
Table 2.
Gene similarity analysis on the basis exon-intron structure between Citrullus lanatus and Arabidopsis thaliana.
Discussion
More studies have focused on the improvement of nutritional quality and tolerance against biotic and abiotic stresses in watermelon (Dabirian et al., 2017; Li et al., 2017; Zhang et al., 2020). However, very few studies have been reported on the seed germination of watermelon. There are different parameters for calculating seed germination such as germination energy, germination rate, vitality index, and germination index. The stored watermelon seeds have a low germination rate, with a specific structure and irregular emergence. For commercial agricultural production, a good and uniform germination rate of watermelon is required. Therefore, it is necessary to improve the germination rate and nutritional quality of watermelon. To improve the seed germination rate, some physical treatments were performed to the seeds like removal of seed coat, scarification, and seed nicking. Few reports exist on the application of chemical substances such as plasma activated water, salicylic acid and oxygenated brackish water which help improve the germination rate in other crops wheat (Triticum aestivum L.), sunflower (Helianthus annuus L.), mung bean (Vigna radiata) and alfalfa (Medicago sativa) (Zhu et al., 2021) to break the seed dormancy.
Seed dormancy is also an important process, like seed germination. The level of dormancy should be well balanced to avoid the seeds from pre-harvest sprouting and to maintain the quality of the harvested seeds. During the seed maturation, if the mother plant is exposed to the environmental factors like temperature, light, soil nitrate, these will stimulate the level of dormancy (He et al., 2014). When the seeds are harvested, they will have a high dormancy level, which gradually decreases during dry seed storage or after ripening.
The role of ABA and GA phytohormones in maintaining seed dormancy and germination was reported over 50 years ago. ABA induces the seed dormancy and GA promotes germination. These two hormones act antagonistically to each other’s signaling and biosynthesis pathways (Gazzarrini et al., 2015). In recent years, in addition to ABA and GA, a few more hormones viz., auxin, ethylene, CTK (cytokinin), jasmonate (JA), BR (brassinosteroid) have been reported to be regulate dormancy and germination in plants (Linkies and Leubner-Metzger, 2012). Among them, ethylene hormone plays an important role in reducing the effects of ABA (Gazzarrini et al., 2015; Linkies and Leubner-Metzger, 2012). The ABA and GA level can be balanced between its biosynthesis and catabolism in seed. The biosynthesized genes for ABA are NCEDs, ZEP, AAO and catabolized by CYP707As. Meanwhile, the list of genes involved in the GA biosynthesis are GA3ox, GA13ox, GA20ox and GA2ox gene in GA catabolism respectively (Tuan et al., 2021).
Genetic and molecular level studies have been carried out to understand the regulation of seed dormancy and have led to the discovery of a few genes (Née et al., 2017). They have identified seven QTLs and named them Delay of germination (DOG1, DOG2, DOG3, DOG4, DOG5, DOG6 and DOG7). Based on the two-mapping populations of RILs, tagging and mapping shortlisted three genes (DOG1, DOG3 and DOG6) responsible for germination (Alonso-Blanco et al., 2003). Both the genes, namely Delay of Germination 1 (DOG1) and Reduced Dormancy 5 (RDO5) act as main regulator for dormancy and germination (Xiang et al., 2014). These regulators have a specific expression in the seeds and show dormancy mutant phenotypes. The DOG1 gene has been cloned, characterized in different plant species, and this protein has been studied further through a transgenic approach (Huo et al., 2016). In the present study, one of the DOG1 gene was upregulated in three varieties at different time points. However, the expression level of this gene was significantly high (200-fold change) in the GN3 variety, suggesting its role more in dormancy than germination. Similar result was observed in another study that DOG1 gene combine with ABA to induce seed dormancy and activate the signaling pathways which had a strong correlation with soil temperature and dormancy levels in buried seeds in soil (Nakabayashi et al., 2012). In addition, one more dormancy regulator was identified through molecular cloning and named Sdr4, which is expressed like rice homolog DOG1 (Sugimoto et al., 2010). Another gene namely RDO2 was used in our gene expression study. Even though this gene is expressed in both GN1 and GN3 genotypes, its expression level was gradually increased in the GN1 genotype. But the expression of this gene was slightly elevated in GN3 genotype at one time point and decreased at a later stage. This result suggests that the RDO2 gene is involved in regulating the germination process.
Another important gene is Mother of FT and TFL1 (MFT), belongs to the member of the plant phosphatidyl ethanolamine binding protein (PEBP) family. In the present study, one of the genes in this family, MTF, was upregulated only in the SI genotype, which has a higher germination rate, revealing that this gene could regulate the germination in watermelon. The MFT gene family has been reported to regulate germination in both Arabidopsis and wheat (Vaistij et al., 2013). Even though the function of this gene is not clearly understood, it negatively regulates the ABA signaling (Xi et al., 2010).
Several genes ABA1, ABA2, ABA3, NCED6, ABI3, ABI4, ABI5, MDN1, PER1, CRU3, CYP707A1, and CYP707A2 were reported to be involved in abscisic acid signalling pathway for seed germination or dormancy. ABA2 gene was upregulated against drought stress or ABA application (Cheng et al., 2002). Transgenic plants overexpressing the ABA2 gene showed delayed germination, primary root growth and tolerance against salinity stress in Arabidopsis (Lin et al., 2007). In our current study, we selected three ABA genes for real time expression analysis. The results revealed that ABA1 was expressed specifically only in the GN1 genotype. Similarly, another gene, ABA3, showed upregulation in both GN1 and GN3 genotypes. However, the expression of this gene gradually increased in the GN1 genotype, while it slowly decreased in the GN3 genotype. In contrast, the ABA2 gene was upregulated ≥ 2 fold in all the genotypes at three different time points. These results suggested that ABA1 and ABA3 genes were involved inducing germination, and ABA2 was involved in both germination and dormancy.
The Abscisic acid insensitive (ABI3) gene belongs to the B3 domain transcriptional factor family, which regulates many genes involved in seed development (Stone et al., 2001). In the present study, one of the ABI3 genes was upregulated only in the SI and GN1 genotypes, suggesting that this gene regulates germination in plants. Similarly, the ABI4 gene encodes an AP2/ERF transcription factor, which acts as an enhancer in the ABA signal pathways in the seed development and germination (Söderman et al., 2000). However, the expression level of this gene is relatively low at almost all stages except during seed maturity and germination (Söderman et al., 2000). This trend of gene expression was also observed in our current study. This gene was expressed in SI and GN1 genotypes having 96 and 54 percent germination, respectively, indicating that it can regulate seed germination. These results suggest that ABI4 plays an important role in balancing the ABA/GA to control the primary seed dormancy. Third gene belong to this group, ABI5 belong to the bZIP transcriptional factor expressed in SI genotype. All the three ABI genes were found to be involved in the seed germination and development.
In our study, we selected two cytochrome P450 genes and validated their expression through real time PCR. Among the two genes, the CYP707A2 gene was expressed uniformly in both SI and GN1 genotypes. Even though, the CYP707A1 gene was expressed in all the three genotypes, higher expression was noted in the GN3 genotype, suggesting that both genes act antagonistically against each other in dormancy and germination. The expression level of cytochrome p450 genes, particularly the CYP707A2 gene, was highly induced in seed during imbibition (Kushiro et al., 2004). Similarly, CYP707A1 and CYP707A2 genes that prevents the accumulation of ABA in seeds showed upregulation after SAEW treatment in watermelon (Wu et al., 2022). Based on the results obtained from ChIP-qPCR and expression assays, ABI4 genes certainly inhibits the CYP707A1 and CYP707A2 genes by directly binding to the CCAC cis-elements that are essential for repression.
In Arabidopsis, the NCED6 gene is expressed only in immature endosperm. This group of genes, NCED6 was upregulated in the GN1 genotype in the present study. The mutant nced6 gene showed reduced ABA levels in dry seeds (Lefebvre et al., 2006). Another regulator namely FUSCA3 (FUS3), controls the mid- and late seed development (Raz et al., 2001). In the present study, one of these genes FUS3 was induced only in the GN1 genotype, which has a 54 percent germination rate. The mutant of this gene shares common phenotype with reduced protein expression in storage seed and reduced dormancy at maturation (Gutierrez et al., 2007). FUS3 gene mutant alleles reported to have a truncated protein and express pleiotropic effects (Tiedemann et al., 2008). The B3 domain gene promoter analysis revealed RY motifs were predicted only in this gene (Braybrook et al., 2006). Cruciferin (CRU) genes are present in the storage proteins in seeds, these CRU1 and CRU2 are bound by ABI3 (Mönke et al., 2012). This family gene, CRU3 was upregulated specifically in the GN1 genotype. During germination, this CRU3 gene is broken down and provide nitrogen sources for the seedlings. Another antioxidant gene, Cysteine Peroxiredoxin 1 (PER1) showed specific expression in seed germination in response to exogenous substances like nitrate, salt and glucose in Arabidopsis (Haslekås et al., 2003). This gene, PER1, was expressed only in the GN1 genotype, suggesting that it induces germination in the seeds.
Dormancy loss is found to be associated with increasing GA receptors GIBBERELLIN INSENSITIVE DWARF1 (GID1a, GID1b and GID1c) involved in the seed imbibition. GA-GID1 association stimulates GA responses through the targeted degradation of negative regulators named DELLA proteins with the motif (Asp-Glu-Leu-Leu-Ala) (Ueguchi-Tanaka et al., 2005). GID1 gene expression was up-regulated during dormancy loss. Similar results were also observed in our study, one of these genes GID1 was induced only in the SI genotype having 96 percent germination rate. During the dormancy loss, to examine the regulation of GID1 expression, ga1-3 mutant was used as a tool for GA biosynthesis. After cold treatment, GID1a, GID1b and GID1c genes were up-regulated, whereas only GID1b gene increased after the ripening. We also selected few more genes reported to regulate the GA signaling pathway they are SCL3, PKL, HD2B, CHR12, GA20x2, and GA20x3 for the gene expression studies. Among these genes, four genes SCL3, PKL, HD2B, and CHR12 genes were up-regulated in both SI and GN1 genotype at different time points, suggesting that these genes may regulate the GA signaling and involve in germination. The other two genes reported to be involved in the GA biosynthesis pathways, GA20x2 gene did not show enhanced levels of activity in any of the genotypes. In contrast, the other gene GA20x3 expressed in all three genotypes. However, the level of expression was seen at two to three time points in SI and GN1 genotypes and slight induction was observed only in GN3 at 12hai. The cold treatments used for seed germination also stimulate the GA biosynthesis genes GA3ox1 and GA3ox2 (Yamauchi et al., 2004). Iglesias-Fernandez and Matilla (2009) reported that SoACS7 and SoACO2 genes inhibits its expression during the early seed incubation but stimulates with the SoGA20ox2, SoGA3ox2, and SoGA2ox6 genes involved in GA metabolism in hedge mustard.
During the germination process, ethylene production increased along with the increase in ACO activity in Arabidopsis and cress (Linkies and Leubner-Metzger, 2012). In contrast, various dormancy breaking treatments like chilling, Gas NO, and HCN also increase ethylene production (Arc et al., 2013). Ethylene acts antagonistic against ABA in germination of various plant species such as Amaranthuscaudatus,Chenopodiumalbum, cotton, tobacco (Nicotianatabacum), and Arabidopsis (Leubner-Metzger et al.,1998). In Arabidopsis, ACO transcripts were upregulated in the aba2 mutant through microarray analysis (Cheng et al., 2009). In the present study, ACO3 was expressed only in the GN1 genotype. In contrast, the ACO4 gene was expressed in all the three genotypes at one time point. These results suggest that ACO3 gene negatively regulates the ABA and ACO4 positively regulates the ABA levels.
miRNAs have been reported to cleave and down regulate the target genes at the post transcriptional level. Auxin Response Factor10 (ARF10) was regulated by microRNA (miRNA) miR160 (Lin et al., 2007). To prove this, transgenic seeds were developed with an miR160-resistant form of ARF10 (mARF10), resulting in sensitivity to germination. In contrast, ectopic expression of miR160 resulted in reduced sensitivity to the ABA treatment (Lin et al., 2007). In the present study, these miRNA156 (PER1, MDN1, CYP707A1) and miRNA390 (MFT, CYP707A2) were present in the multiple genes. The miRNA166 is unique to ABI5 gene, miRNA167 to GA20x3. The miRNA156, miRNA390, miRNA166, miRNA167 is reported to be involved in early stages of seed germination in Brassicanapus (Jian et al., 2016). Another miRNA396 is also detected in multiple genes namely CHR12, ABI5, CYP707A1 and CYP707A2 in our study. The miRNA396 acts as a growth regulating factor genes and regulate the expression against beet cyst nematodes in Arabidopsis (Hewezi and Baum, 2012).
The promoter analysis of ABA2 gene (At1g52340) revealed three ABRE like elements (TACGTGTC, TACGTG, and GCCACGTGT). In AtNCED3 (At3g14440), revealed one dehydration response element (TACCGACAT) and two ABRE like elements (TACGTC, TACGTG, and GACCGTA) were identified, suggesting its role in drought stress (Lin et al., 2007). In the present study, we also detected ABRE like elements in most of the genes, suggesting their importance in seed germination or dormancy. We have also detected some specific cis-elements like RY-elements in the CRU3 gene in our study. A similar element was also observed in other genes, B3 domain gene promoter analysis revealed RY motifs specific to this gene (Braybrook et al., 2006). The CYP707A1 and CYP707A2 genes by directly binding to the CCAC cis-elements that are essential for repression. The ABI4 gene is a multipurpose transcription factor that binds to the CCAC element to inhibit the transcription of some genes (Bossi et al., 2009). In our study, we have observed an AACA motif present in the promoter anlaysis of the ABI3 gene. More detailed studies should be conducted further to understand its exact role in seed germination or dormancy.
Our study is the first attempt to understand the genes involved in seed germination and seed dormancy of watermelon through expression studies. We have selected genes reported to be involved in different pathways like GA, ABA, ethylene, jasmonate, auxin, etc., and analyzed their expression in the watermelon lines. Based on the results, two genes, DOG1 and CYP707A1 could be involved in the seed dormancy. However, Pickle (PKL), ABI3, ABI4, MDN1, SCL3, and CHR12 genes were significantly involved in the seed germination process. In addition, some miRNAs viz., miR-156, miR-396, miR-171, and miR-390 were predicted in multiple genes. However, the exact role of these miRNAs can only be clearly explained after additional experiments are carried out.