Introduction
Materials and Methods
Sample Collection
DNA Extraction and PCR
Analysis
Result
Taxonomic Relationship of Epilobium in Haenam Population
Genetic diversity
Analysis of genetic differentiation
Phylogenetic relationships
Discussion
Introduction
Epilobium hirsutum L.(Onagraceae) is a perennial herbaceous plant native to Russia, Japan, China, India, and South Korea. The flower solitary in upper axils, pink, stigma deeply 4-cleft (e-Flora of South Africa, 2022), and the fruits is defined as the inferior dorsal-seprifragal tetralocular many- seeded capsule with synascidiate and symplicate zones (Odintsova and Klimovych, 2017). Notably, the characteristic of 4-cleft in the stigma has been identified as a useful trait to distinguish it from other species in the genus (Lee et al., 2013).
The northern plant, E. hirsutum, has confined populations and individuals in the central part of the Korean Peninsula. It is recorded in about five populations in Gyeongsangbuk-do and Gangwon-do (National Institute of Biological Resources, 2012). Particularly, the populations, in wet places near streams, have been destroyed by the road constructions and has been legally designated and protected as Level Ⅱ Endangered Wild Plants (National Institute of Biological Resources, 2012), and classified as a vulnerable (VU) species according to their red list of the International Union for Conservation of Nature (IUCN) by Ministry of Environment.
Intra-specific genetic diversity provides insights into the demographic and evolutionary histories of populations (Paz-VInas et al, 2018). Moreover, It is well known that preserving the genetic diversity of endangered species can significantly affect their long-term survival and evolution in changing environments (Frankham et al., 2002). However, it is difficult to plan conservation strategies for endangered species because of the insufficiency of genetic information, such as genetic diversity, genetic structure, and genetic differentiation among different populations.
Accordingly, we conducted to investigate the genetic diversity and genetic differentiation among populations of E. hirsutum, in order to establish a foundation for the conservation and restoration of this plant population.
Materials and Methods
Sample Collection
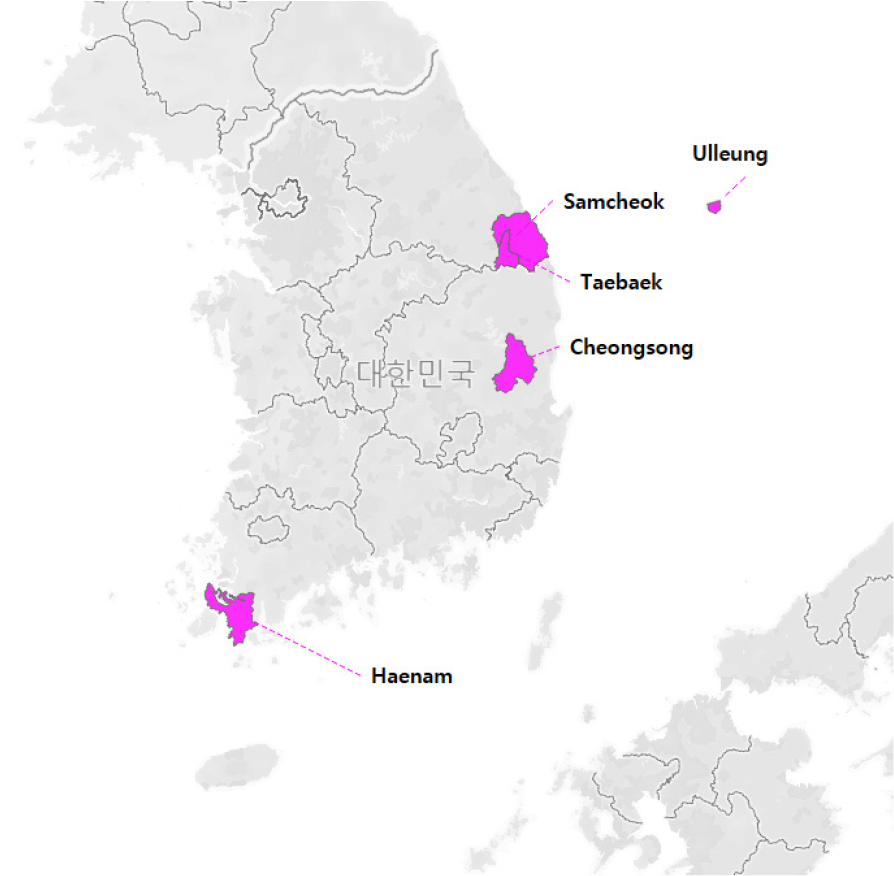
To understand the distribution status of E. hirsutum in South Korea in detail, specimen and reference were collected. As a result, it was confirmed that E. hirsutum is extremely rare in areas, such as Samcheok-si and Taebaek-si in Gangwon-do, as well as Cheongsong-gun and Ulleung-gun in Gyeongsangbuk-do (National Institute of Biological Resources, 2012). The samples for this study were collected from a total of 75 individuals in four population. Additionally, a total of 17 individuals were collected from Haenam-gun, which has been reported as a new habitat (Fig. 1, Table 1). Samples were collected after obtaining collection permits for endangered wildlife from the respective regional environmental offices, as E. hirsutum is classified as an Endangered species Ⅱ. To represent each population while considering their ecological characteristics and geographical distribution, only mature individuals that were at least 1 meter tall and had developed reproductive organs were selected, with a minimum distance of 1 meter maintained between individuals. Additionally, to minimize damage to the natural habitat, only one leaf per individual was collected for DNA extraction.
Table 1.
Sampling information of Epilobium hirsutum L.
DNA Extraction and PCR
The DNA extraction followed the method that was reported in Loockerman and Jansen (1996) modified CTAB method of Doyle and Doyle (1987). primers used in this study were mlh1 and matK. PCR was performed in 25 µL, containing template DNA (20~100ng), 10X Diastar™ Taq DNA buffer (2.5 μL, Solgent Co.), 10mM dNTPs mix (0.5 μL, Solgent Co.), Forward primer (10 pmol), Reverse primer (10 pmol), and Diastar™ Taq DNA polymerase (0.12 μL, Solgent Co.). The amplification procedure was pre-denaturation at 95°C for 5 min, followed denaturation by 35 cycles at 95°C for 50s, 50~60°C for 40s (each pair of primer was screened in gradient temperature), extension at 72°C for 1 min, and final extension at 72°C for 10 min. PCR product was determined by electrophoresis in 1.5% agarose gel.
Analysis
All amplicons were sequenced using the ABI PRISM BigDye® Terminator v3.1 Cycle Sequencing Kit (PE Applied Biosystems, USA). De novo assembly of the paired-end raw reads was performed using Geneious R11 software (Biomatters, 2017).
To assess the phylogenetic position of the Haenam population, a set of matK sequences from related taxa was downloaded from NCBI GenBank (Table 2). A maximum likelihood phylogenetic tree was then reconstructed in IQ-TREE using the GTR+G substitution model and 1,000 bootstrap replicates.
Table 2.
List of matK sequences of related taxa retrieved from NCBI GenBank.
For the mlh1 gene, DnaSP v6 (Rozas et al., 2017) was used to compute genetic diversity indices, including nucleotide diversity (π), haplotype diversity (h), Shannon’s information index, the number of polymorphic loci, and Nei’s genetic distance. A dendrogram was constructed using the unweighted pair group method with arithmetic mean (UPGMA). In addition, analysis of molecular variance (AMOVA) based on genetic distance was conducted using Arlequin v3.0 (Excoffier et al., 2005).
Result
Taxonomic Relationship of Epilobium in Haenam Population
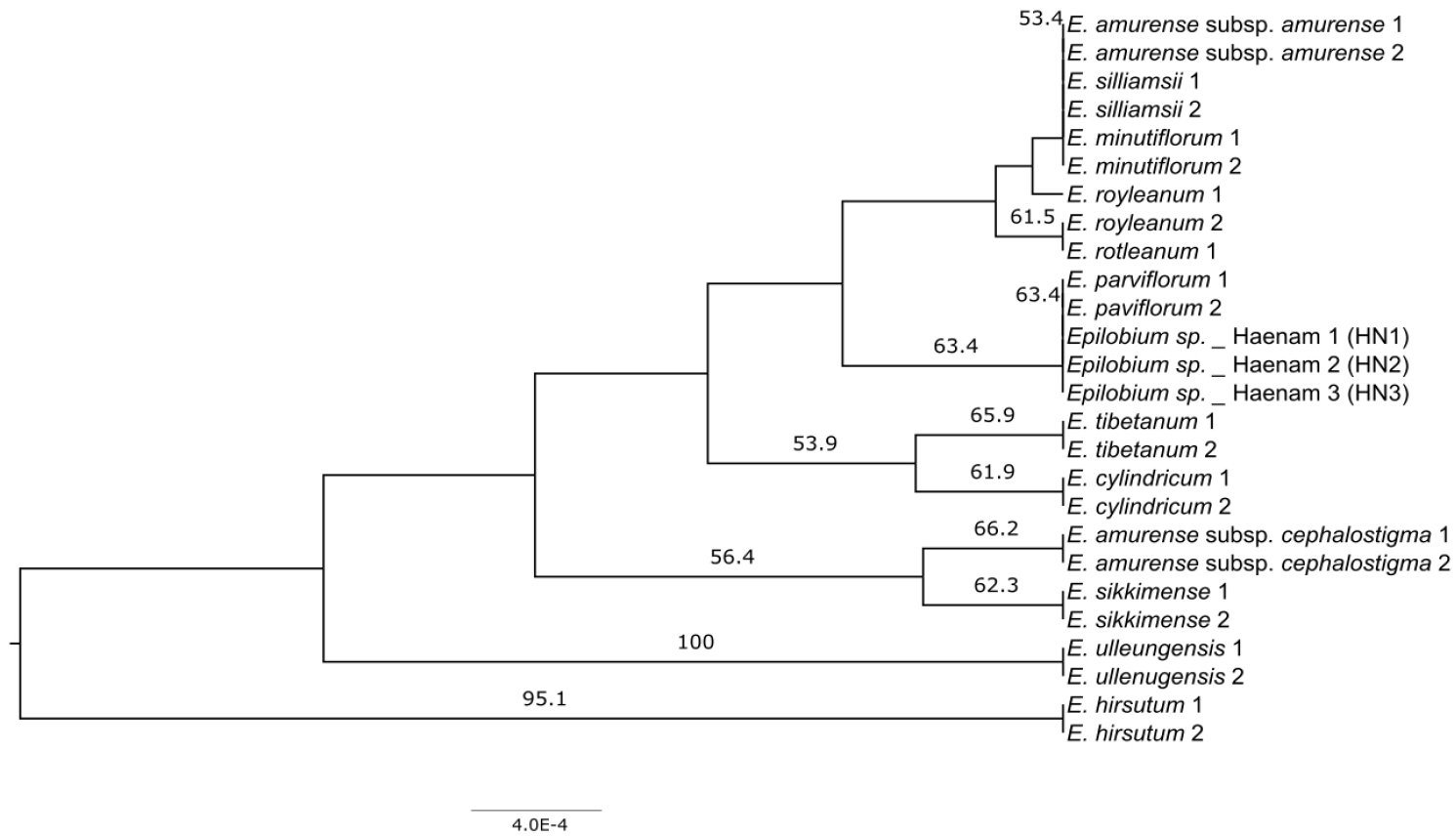
Compared the matK sequences of 11 taxa of the genus Epiloibum available on NCBI showed variation at 9 SNP sites (Table 3). Among them, E. hirsutum showed autapomorphy at 3 sites (241bp, 470bp, 733bp), while the Haenam (HN) population showed a synapomorphy at 1 site (139 bp) shared with E. paviflorum, indicating a clear distinction from other populations. To confirm the taxonomic status of the HN population, we conducted a neighbor-joining (NJ) tree, with only those clades displaying bootstrap values of 50% or higher indicated on the tree. The NJ tree showed that the HN population is a monophyletic group with E. paviflorum, with a bootstrap value of 63.4% (Fig. 2).
Table 3.
DNA sequence variation of matK among Epilobium L. in Korea.
Genetic diversity
To analyze the genetic diversity of E. hirsutum, a total sequence length of 238 bp from the mlh1 gene region was evaluated to assess the genotypes (haplotypes) of 4 populations. A total of 4 haplotypes (H1–H4) were identified, and 4 regions of nucleotide variation were confirmed (Table 4). H1 and H2 is observed in the SC population, H2 in the CS population, H2 and H3 in the TB population, H4 in the UL population.
Table 4.
Haplotype analysis of the mlh1 gene.
| Haplotype | 1 | 22 | 34 | 353 |
| H1 | G | G | C | A |
| H2 | T | G | C | A |
| H3 | T | G | T | A |
| H4 | T | A | T | C |
Moreover, genetic diversity indices varied across populations of E. hirsutum, as listed in Table 5. Haplotype diversity (Hd) ranged from 0.0000 to 0.47619, with the UL and CS populations having the lowest (0.00) and the SC population having the highest (0.47619). Similarly, nucleotide diversity (π) also ranged from 0,000 to 0.00412. We performed the Tajima’s D and Fu’s Fs tests to detect past population evolution. Tajima’s D values is negative for the TB population (-1.13197) but statistically not significant. Additionally, Fu’ s F-value of all population ranged positively from 0 to 0.952.
Table 5.
Genetic diversity information of the four populations of E. hirsutum.
| ID | hz | Hdz | Nucleotide diversityz | Dz | Fsz |
| SC | 2 | 0.47619 | 0.00294 | 0.55902 | 0.589 |
| TB | 2 | 0.33333 | 0.00412 | -1.13197 | 0.952 |
| CS | 1 | 0 | 0 | 0 | 0 |
| UL | 1 | 0 | 0 | 0 | 0 |
Analysis of genetic differentiation
Analysis of genetic differentiation among the four populations of E. hirsutum revealed that Pairwise-FST ranged from 0.00037 to 0.64159, indicating significant genetic differentiation among all populations (Table 6). The SC and TB regions had the closest genetic distance with an FST value of 0.00037. In contrast, the geographically isolated UL and CS population showed an FST value of 0.64159, indicating that the two populations share a little genetic diversity.
Table 6.
Genetic differentiation among population (Pairwise-FST).
| Population | SCz | TBz | CSz | ULz |
| SC | - | 0.00037 | 0.06816 | 0.34928 |
| TB | 0.00037 | - | 0.10562 | 0.15112 |
| CS | 0.06816 | 0.10562 | - | 0.64159 |
| UL | 0.34928 | 0.15112 | 0.64159 | - |
Phylogenetic relationships
AMOVA (Analysis of molecular variance) results showed that 84.7% of the total genetic variation was due to differences within populations, while the remainder (15.3%) occurred among regions (Table 7). Based on Nei’ s genetic distance values among populations, the dendrogram was constructed using UPGMA, and indicated that the distribution of E. hirsutum tends to cluster with UL and TB, with SC and CS clustering separately, followed by the HN population (Fig. 3).
Table 7.
Analysis of molecular variance (AMOVA) between four population of E. hirsutum.
| Source of variation | Sum of squares | Variance components | Percentage variation |
| Among populations | 25.754 | 0.03801 | 15.26 |
| Within populations | 143.036 | 8.4387 | 84.74 |
| Total | 168.79 | 8.47671 |
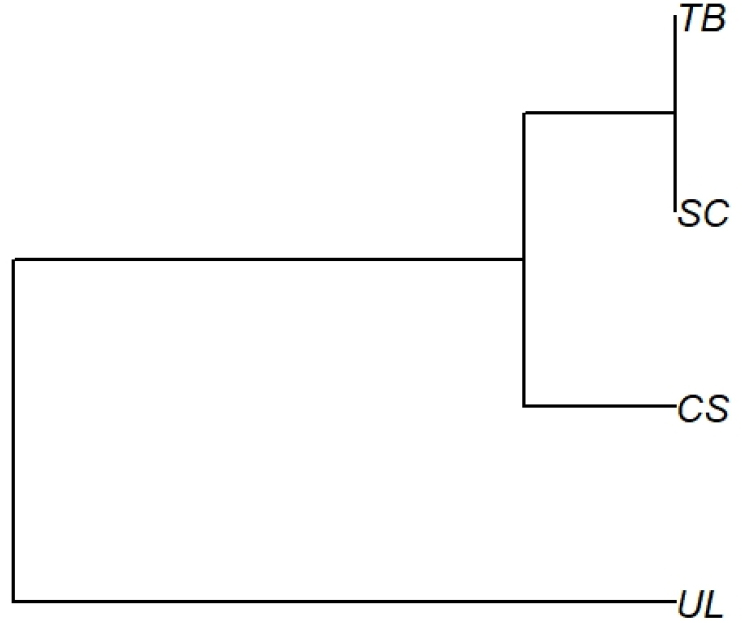
Fig. 3.
Dendrogram using the UPGMA method based on Nei (1978) genetic distance five population of Epilobium hirsutum L.
Discussion
The results of our study provide insights into the genetic diversity of Epilobium hirsutum based on the nuclear mlh1 gene, and clarify the taxonomic identity of the Haenam (HN) population using the chloroplast matK gene, which represents a recently reported habitat.
E. hirsutum, a northern temperate species, is extremely rare in South Korea, being confined to isolated habitats in Samcheok and Taebaek (Gangwon-do), and Cheongsong and Ulleung-do (Gyeongsangbuk-do) (National Institute of Biological Resources, 2013). The Haenam population, recently reported from the southern region of Haenam, is morphologically similar to E. hirsutum, but this identification had not been confirmed. To clarify its taxonomic position, we performed a preliminary phylogenetic analysis using the matK gene. The resulting phylogenetic tree showed that the Haenam population formed a distinct monophyletic clade with E. parviflorum, supported by a high bootstrap value and a 4bp sequence difference from E. hirsutum. These results suggest that the HN population may be more accurately classified as E. parviflorum rather than E. hirsutum.
Previous molecular studies targeting nrITS, matK, and the rps16 intron failed to detect sequence variation among Korean E. hirsutum populations (National Institute of Biological Resources, 2013), emphasizing the need for more informative nuclear markers. Nuclear genes have recently been adopted for phylogenetic studies of angiosperms due to their biparental inheritance and faster evolutionary rates, which provide better resolution among closely related species (Elizabeth et al., 2012). However, orthology must be carefully evaluated, as nuclear genomes often exhibit hybridization and polyploidy, adding complexity to phylogenetic inference (Soltis et al., 2009). In this study, the nuclear mlh1 gene provided useful resolution for assessing genetic diversity and population structure.
The overall nucleotide diversity across the E. hirsutum populations ranged from 0 to 0.00412, indicating a low level of genetic variation. Such limited diversity suggests that these populations, particularly those at the species’ southern edge or in relict habitats, are vulnerable to environmental change and may face increased extinction risk under stochastic events (Guo et al., 2023).
Notably, both the CS and UL populations showed no genetic variation at the mlh1 locus, with haplotype and nucleotide diversity values equal to 0. This may indicate recent bottlenecks or founder effects, which can elevate the risk of extinction due to reduced adaptive potential. Populations with such low genetic diversity should be prioritized for conservation, through both in situ monitoring and ex situ genetic safeguarding strategies.
The AMOVA results confirmed the presence of genetic structure. Molecular variance was partitioned into 84.7% within populations and 15.3% among populations. This pattern, consistent with previous studies on outcrossing plant species (National Institute of Biological Resources, 2014), suggests that E. hirsutum maintains substantial within-population diversity, despite fragmented distributions. To further investigate demographic history, we applied neutrality tests including Tajima’s D (Tajima, 1989) and Fu’s Fs. Both values were negative but not statistically significant, indicating no strong evidence for recent population expansion or purifying selection at the mlh1 locus.
Genetic differentiation among populations varied widely, with pairwise Fst values ranging from 0.00037 to 0.64159. The SC and TB populations showed very low differentiation (Fst = 0.00037), reflecting close genetic similarity and suggesting potential ongoing gene flow due to their geographic proximity. In contrast, the UL and CS populations exhibited high genetic differentiation, likely due to historical isolation. In particular, the geographic isolation of Ulleung-do, as an island, may have contributed to its genetic distinctiveness. These results demonstrate the presence of diverse genetic structures among the sampled populations. It should also be noted that E. ulleungensis, an endemic species newly described in 2017 from Ulleung-do, has been proposed to be of hybrid origin, likely involving E. hirsutum and either E. palustre or E. tetragonum (Chung et al., 2017). This raises the possibility that the Ulleung population analyzed in our study may not be pure E. hirsutum, but rather a cryptic hybrid lineage. This taxonomic uncertainty complicates the interpretation of genetic diversity results for the UL population and underscores the need for further genomic studies to elucidate its evolutionary history.
Accelerated global warming caused by climate change is increasingly exerting direct and indirect impacts on forest ecosystems worldwide, leading to a decline in plant species richness and structural simplification (Jump and Peñuelas, 2005; Kim et al., 2024). In this context, the conservation of rare and endangered wetland species such as E. hirsutum is becoming increasingly critical. Preserving genetic diversity and evolutionary potential is a fundamental goal in species management and recovery planning (Margules and Pressey, 2000). In Korea, E. hirsutum is primarily found in valleys and wetlands, which are fragmented and vulnerable to anthropogenic disturbances such as river maintenance and road construction. The study areas Taebaek, Samcheok, Cheongsong, and Ulleung are all exposed to similar threats of habitat degradation due to anthropogenic disturbances such as development. However, site-specific disturbance factors were also identified, indicating the need for tailored conservation strategies. In Taebaek, environmental improvement measures are required to address pollution originating from nearby agricultural fields and livestock farms. In Cheongsong, the native population is under severe pressure from the invasive species Humulus japonicus, necessitating active management such as the removal of this ecological threat. In Ulleung, large-scale construction activities have already caused significant damage to the natural habitat and population. Moreover, due to its exposed environment, the area faces additional risks such as illegal harvesting and habitat destruction, making the population highly vulnerable to imminent extinction. Therefore, urgent and proactive population management measures are essential to mitigate the impact of human disturbances. Given the accelerating degradation of these habitats and population decline, both in situ and ex situ conservation strategies must be urgently implemented (Wang, 2020).